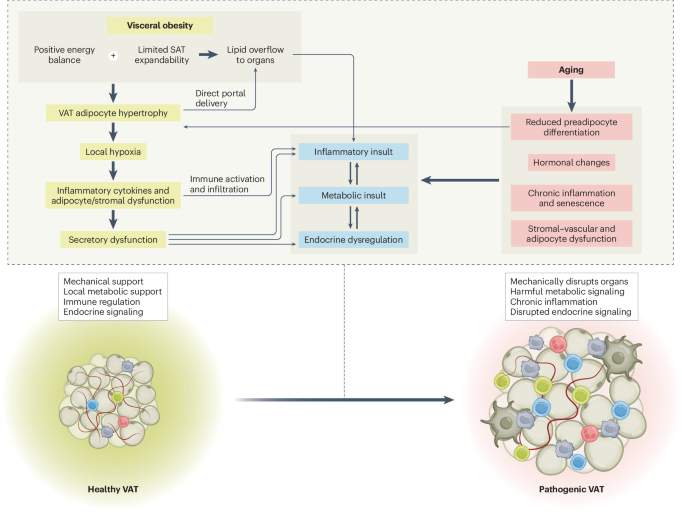
Maternal diet-induced alterations in uterine fluid sncRNAs compromise preimplantation embryo development and offspring metabolic health
Animals
All experimental procedures involving Crl:CD1 (ICR) mice (Charles River Laboratories China) were conducted in strict compliance with protocols approved by the Institutional Animal Care and Use Committee (IACUC) of Beijing Normal University (CLS-EAW-2021-005). The study adhered to established guidelines aimed at minimizing animal suffering. 8-week-old mice were housed under standard laboratory conditions with unrestricted access to food and water, maintained on a 12:12-h light/dark cycle at 22–25 °C and 40–60% relative humidity. Eight-week-old female ICR mice were co-housed with male ICR mice, and pregnancy was confirmed by the presence of a vaginal plug, designated as embryonic day 1 (day 1). Pregnant females were then housed individually. In the HFD group, pregnant females were fed a high-fat diet (HFD, 60% fat, 20% protein and 20% Carbohydrate, D12492, Research Diets Inc.), or normal diet (ND, 5% fat, 17% protein, Beijing HFK Bioscience Co) from day 1 to day 4 of pregnancy, after which they were switched to ND starting on day 5 and maintained on ND until parturition. The control group received a normal diet throughout the entire pregnancy. Blastocysts of both groups were collected on D4 from the uteri for transcriptomic sequencing analysis. On day 5, implantation sites were identified by visualizing blue-stained regions in the uterus following intravenous injection of 0.1 mL of 0.1% trypan blue. On day 12, pregnant mice were humanely euthanized, and their uteri were harvested, fixed overnight in Bouin’s solution, and weighed. Litter sizes and neonate weights were recorded at birth, and fetal lengths were measured.
Glucose tolerance test (GTT)
GTT was performed on dams on day 4 of pregnancy, which were fed either ND or HFD, as well as on their offspring at 2, 4, and 6 months of age. Both male and female progeny were included in the analysis. For the GTT, mice were fasted for 12 h prior to testing. Baseline blood glucose levels were measured from tail vein samples using a glucometer. Mice were then administered glucose intraperitoneally at a dose of 1.5 g/kg body weight. Blood glucose levels were subsequently measured at 0, 30, 60, 90, and 120 min following glucose administration. Data were presented as line graphs, and the area under the curve (AUC) was calculated to assess total glucose exposure over time.
Oviductal fluid and uterine fluid collection
Oviductal fluid (OF) was collected at 18:00 on days 1–3, and UF was collected at 18:00 on day 4, using methods adapted from previous studies32. Briefly, after anesthetizing the mice, uteri or oviducts were excised, and carefully trimmed of excess mesometrium and adipose tissue, rinsed twice in pre-cooled Dulbecco’s Phosphate-Buffered Saline (DPBS. RNase-free), and blotted dry with filter paper. For UF collection, the uterine horns were incised, and cold DPBS was gently injected through the cervical end using a 1 mL syringe. OF was collected by flushing cold DPBS through the oviductal opening. Collected fluids were transferred to sterile glass dishes, where embryos were carefully removed under a stereomicroscope using a mouth pipette. The fluids were then transferred to 1.5 mL centrifuge tubes and centrifuged at 300 × g for 20 min at 4 °C to remove tissue debris, followed by a second centrifugation at 1000 × g for 20 min at 4 °C to eliminate blood cells and other impurities. The supernatant was mixed with three volumes of TRIzol LS reagent, vortexed vigorously, incubated at room temperature for 5 min, and stored at −80 °C for subsequent analysis.
Isolation of 15–50 nt small RNAs
Total RNA from oviductal and uterine fluids was extracted using the TRIzol LS protocol. The mixture was thoroughly vortexed, incubated at room temperature for 15 min, and phase-separated by adding 200 μL of chloroform per mL of solution, followed by vigorous shaking for 10 s and incubation at room temperature for 10 min. Samples were centrifuged at 12,000 × g for 15 min at 4 °C, and the upper aqueous phase (~500 μL) was carefully transferred to a new tube. RNA was precipitated by adding an equal volume of isopropanol and 1 μL of glycogen, followed by gentle inversion and overnight incubation at −20 °C. The RNA pellet was obtained by centrifugation at 12,000 × g for 15 min at 4 °C, washed twice with 1 mL of 75% ethanol, air-dried, and resuspended in 10 μL of RNase-free water. RNA concentration and purity were measured using a NanoDrop 2000C spectrophotometer. RNA fragments ranging from 15 to 50 nucleotides (nt) were isolated using a 10% denaturing polyacrylamide gel, following a protocol adapted from our previous work29. Total RNA was electrophoresed on a 10% urea-polyacrylamide gel at 200 V for 40 min in 1× Tris-Borate-EDTA (TBE) buffer (Thermo Fisher Scientific). The gel was stained with SYBR Gold nucleic acid gel stain (Thermo Fisher Scientific), visualized using a UV transilluminator, and RNA fragments within the 15–50 nt range were excised using RNA molecular weight markers (New England Biolabs) as guides.
Quantitative analysis of RNA modifications in mouse uterine fluid using LC-MS/MS
Uterine fluid samples were collected from pregnant mice at gestation day 4, with three biological replicates analyzed per dietary group (ND and HFD). Each biological replicate comprised pooled samples from four mice to ensure sufficient samples. 15–50 nt RNA isolated from mouse uterine fluid was enzymatically digested into mononucleotides in a 50 μL reaction mixture containing 5 μL of 10× reaction buffer (2.5 M Tris-HCl, pH 8.0, 50 mM MgCl2, 5 mg/mL bovine serum albumin [BSA]), 1 IU Benzonase (Sigma-Aldrich), 0.2 IU alkaline phosphatase (Sigma-Aldrich), 0.01 IU phosphodiesterase I (USB), and nuclease-free water, and incubated at 37 °C for 3 h. The digest was filtered using a Nanosep 3 K device with Omega membrane (Sigma-Aldrich) at 14,000 rpm for 20 min at 4 °C, then the flow-through was transferred to a chromatographic vial for LC-MS/MS analysis as described previously71,72. Nucleoside standards (cytidine, adenosine, guanosine, uridine, m¹A, m6A, Am, I, Im, hm5C, m5C, ac4C, Cm, m¹G, m2G, m7G, m²,²G, Gm, Um, m3U, m5U, m5Um, and Ψ) served as quantification controls, with calibration curves generated from a gradient dilution series. Replications were individually processed and subjected to single-injection analysis. Chromatography was performed on an Agilent 1200 series LC, using a 2.1 mm × 150 mm Hypersil GOLD aQ C18 column, 3 µm particles (Thermo Fisher) held at 40 °C. MS analysis was performed by a triple quadrupole mass spectrometer (ThermoFisher TSQ Vantage) with an electrospray ionization source in a positive ion mode using multiple reaction monitoring. Raw data from LC-MS/MS were acquired and subsequently processed with Thermo Xcalibur 3.1 data system. Absolute amounts of each ribonucleoside were back-calculated from the contemporaneous calibration curve. To control for variation in sample loading, the molar concentration of every modified ribonucleoside was normalized to the total amount of nucleosides sharing the same nucleobase (e.g., m5C% = m5C/[m5C + Cm + hm5C + ac4C + C]).
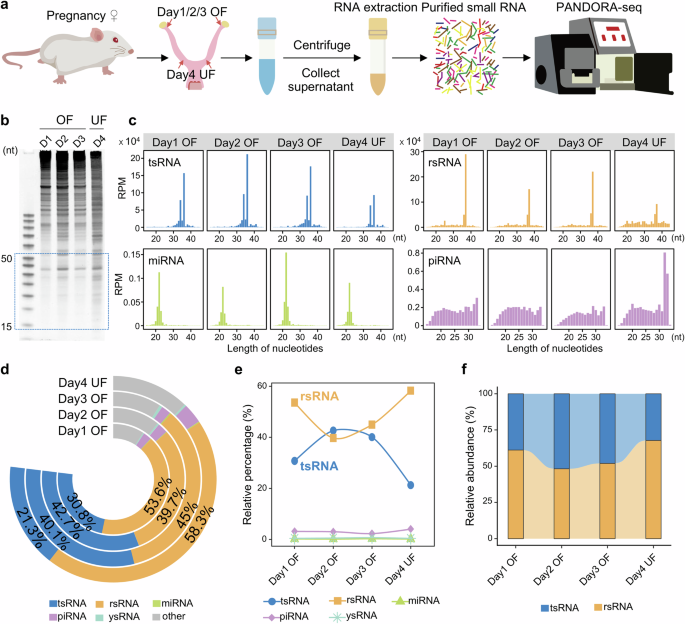
Detection of small RNA expression patterns using PANDORA-seq
Small RNAs were treated following the PANDORA-seq protocol, as previously described29,30. Small RNAs were treated with T4 polynucleotide kinase (T4PNK) reaction mixture (5 μL 10 × PNK buffer, 1 mM ATP, 10 U T4PNK) followed by RNA isolation. The collected RNAs were then treated with α-ketoglutarate-dependent hydroxylase (AlkB) mixture (50 mM HEPES, 75 μM ferrous ammonium sulfate, 1 mM α-ketoglutaric acid, 2 mM sodium ascorbate, 50 mg/L BSA, 4 μg/mL AlkB, 2,000 U/mL RNase inhibitor) followed by RNA isolation with TRIzol LS. Then the small RNA libraries were constructed and sequenced using DNBSEQ platform (BGI Genomics Co., Ltd., Shenzhen, China) with 50-bp single-end reads.
Then small RNA sequences were annotated using the software SPORTS1.173 with a one-mismatch tolerance. Reads were then mapped to several non-coding RNA databases, including: (1) miRNA database miRBase 22.1; (2) genomic tRNA database GtRNAdb (mm39 for mouse); (3) mitochondrial tRNA database mitotRNAdb; (4) rRNA and YRNA databases assembled from the National Center for Biotechnology Information nucleotide and gene database; (5) piRNA database piRBase v3.0; and (6) non-coding RNAs defined by Ensembl and Rfam 14.7.
Differential expression analysis was conducted using the R package DESeq2, identifying significantly differentially expressed small RNAs (defined based on an average RPM > 1) with a threshold of adjusted P ≤ 0.05 and an absolute |log2 fold change| ≥ 1.
Selection of sncRNAs for transfection
Differential expression analysis was conducted to identify sncRNAs significantly upregulated in the HFD group relative to the ND group, focusing on four major classes defined by molecular origin and annotation: genomic tRNA-derived small RNA (cyto-tsRNA), rRNA-derived small RNAs (rsRNAs), mitochondrial tRNA-derived small RNAs (mt-tsRNAs), and microRNAs (miRNAs). Candidate miRNAs were selected based on prior reports of functional relevance in metabolic regulation. To refine tsRNA and rsRNA selection, class-specific abundance thresholds were applied, retaining sequences with log2FC > 1, baseMean > 500 and adjusted P ≤ 0.01. Only two mt-tsRNAs meet the criterion, which were chosen. As cyto-tsRNAs predominantly originated from Leu, Trp, His, Glu, and Ser tsRNAs, while rsRNAs were mainly derived from 18S and 28S rRNAs, we selected the top sequences from tsRNA-Leu, tsRNA-Trp, tsRNA-His, tsRNA-Glu, tsRNA-Ser, mt-tsRNA, 18S-rsRNA, 28S-rsRNA separately, by considering the lower adjusted P, higher log2FC, higher baseMean, smaller lfcSE (the standard error estimate for the log2 fold change estimate) and lower sem (standard error of the mean).
Transfection of murine morulae with sncRNAs
On the morning of day 3, morulae were isolated from mouse oviducts and subjected to four washes in M2 medium. The embryos were then transferred into acidic Tyrode’s solution in a covered dish, ensuring that they remained suspended near the solution’s surface throughout the process. The zona pellucida was dissolved within 15–30 s under continuous microscopic observation, taking care to prevent embryos from adhering to the dish. Following complete dissolution, embryos were transferred to fresh M2 medium, washed 1–2 times to remove any residual acidic solution, and cultured in KSOM medium. Transfection procedures followed established protocols, with four experimental groups corresponding to different RNA types and an empty liposome control group that underwent the same treatment process but without the addition of sncRNAs. The transfected sncRNA groups included: sncRNA pool, tsRNA pool, rsRNA pool, miRNA pool. Sequences of the sncRNAs used were listed in Supplementary Table 2. Transfections were performed using Lipofectamine™ (STEM00001). Solution A was prepared by mixing 0.5 μL of Lipofectamine™ with 25 μL of KSOM medium, while a 5 nM concentration of 5′ FAM-labeled RNA pool was mixed with 25 μL of KSOM medium to prepare Solution B. Solutions A and B were combined, incubated for 10–15 min to allow complex formation, and then applied in 50 μL droplets covered with mineral oil to prevent evaporation. The droplets were equilibrated in an incubator at 37 °C for at least 30 min before 10 morulae were cultured in one droplet of transfection mixture for 24 h. Transfection efficiency was assessed by fluorescence microscopy, evaluating the localization of the fluorescent signal within the morulae. Following transfection, blastocysts were collected and stored in RNA preservation solution for subsequent sequencing analysis. Each transfection experiment was independently repeated three times.
NIH/3T3 cell transfection and QPCR detection
In the NIH/3T3 cell transfection experiments, NIH/3T3 cells with a passage number below 20 were sourced from the National Cell Resource Center (1101MOU-PUMC000018, Beijing, China). Transfection procedures were carried out according to established protocols. Specifically, the following were transfected into NIH/3T3 cells separately: sncRNA pool, tsRNA pool, rsRNA pool, miRNA pool, individual sncRNAs (tsRNA-Leu-AAG: GGUAGCGUGGCCGAGC, mt-tsRNA-Trp-TCA: AGUCCGCGAGCCUUC, and rsRNA-18S: GAGCGGUCGGCGUCCC), and controls (empty liposome transfection without sncRNAs). Transfection efficiency was evaluated 48 h post-transfection using fluorescence microscopy, and cell proliferation was assessed using the MTT assay. Cells transfected with individual sncRNAs and controls were harvested 48 h after transfection. Total RNA was extracted and reverse-transcribed into cDNA. Quantitative PCR was then performed using the primers, which listed in Supplementary Table 3.
Transcriptome sequencing of blastocysts and data analysis
Total RNA from blastocysts was isolated and enriched for polyadenylated mRNA using Oligo (dT) magnetic beads, followed by reverse transcription and amplification according to the Smart-seq2 protocol. The resulting double-stranded complementary DNA (cDNA) was treated with Tn5 transposase for fragmentation and adapter ligation, constructing libraries suitable for high-throughput sequencing. Library quality was assessed using an Agilent 2100 Bioanalyzer and quantitative PCR (qPCR) before sequencing on the DNBSEQ platform, generating paired-end 100 base pair (PE100) reads, providing a minimum of 6 gigabytes (GB) of high-quality data per sample. Raw sequencing data underwent rigorous quality control using SOAPnuke (v1.5.6), which included removing reads containing adapters, reads with more than 5% unknown bases (N), and low-quality reads where more than 20% of bases had a quality score below 15. Clean data were aligned to the mouse reference genome using Bowtie2 (v2.3.4.3), and gene expression was quantified using RNA-Seq by Expectation-Maximization (RSEM) software (v1.3.1). Principal component analysis (PCA) was performed using the FactoMineR package, with visualizations generated by factoextra. Differentially expressed genes (DEGs) were identified using the DESeq2 package, with significance thresholds set at |log2 fold change| ≥ 1 and adjusted P ≤ 0.05. Volcano plots of differential gene expression were created using ggplot2, and gene expression clustering across samples was visualized using heatmap. Functional annotation and pathway enrichment analyses of DEGs were performed using the clusterProfiler package. The Benjamini-Hochberg method was used to correct for multiple testing. Gene Ontology (GO) term enrichment analyses were conducted, with significance thresholds set at an adjusted P ≤ 0.05.
Statistics and reproducibility
Mice were randomly sorted into ND and HFD groups. Sample sizes were determined based on previous studies and power calculations. Two-tailed Student’s t-tests were conducted to compare the relative abundance of sncRNAs, embryo/placental weight, survival numbers/rates, litter size, newborn birth weights/lengths, GTT results between the ND and HFD groups, gene set scores for GOBP terms between control and RNA pool transfection, as well as relative gene expression levels between control and individual sncRNA transfections. For the analysis of RNA modification dynamics, a two-tailed multiple t-test was performed using GraphPad Prism. Correlations between sncRNAs and genes were assessed using Spearman’s rank correlation analysis to generate the correlation coefficient (ρ). Statistical significance was set at P ≤ 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link