Integrative analysis of gut microbiota and metabolic pathways reveals key microbial and metabolomic alterations in diabetes
Microbial diversity and compositional shifts in diabetic versus control groups
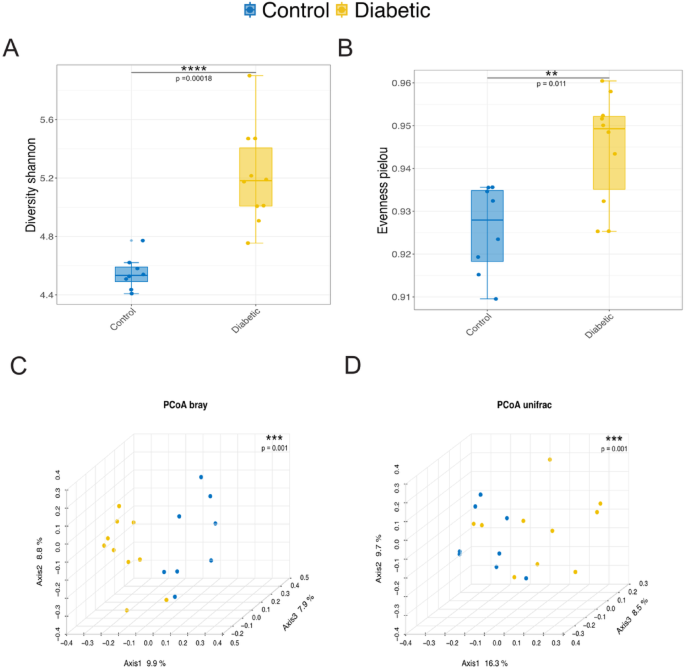
The analysis revealed a higher microbial diversity in the diabetic group compared to the control group, as indicated by the Shannon diversity index (Fig. 1A). Additionally, the diabetic group exhibited a more even distribution of species, with a higher Pielou’s evenness index (Fig. 1B). Conversely, the control group showed a less uniform distribution of species. The Principal Coordinate Analysis (PCoA) based on Bray-Curtis dissimilarity demonstrated distinct separation between the diabetic and control groups, with 9.9% of the total variation (Fig. 1C). Furthermore, the phylogenetic composition of the communities displayed noticeable differences, as evidenced by the PCoA plot based on unweighted UniFrac distances (Fig. 1D). The distinct clustering observed along Axis 1, which accounts for 16.3% of the variation, highlights that the microbial communities in diabetic individuals are compositionally different and phylogenetically distinct from those in the control group. This underscores the impact of diabetes on the gut microbiota’s structure and evolutionary relationships. Furthermore, individuals with diabetes seem to possess a richer and more diverse gut microbiota, suggesting a more balanced microbial ecosystem. These differences reflect significant variations in the relative abundance of microbial species between the two groups.
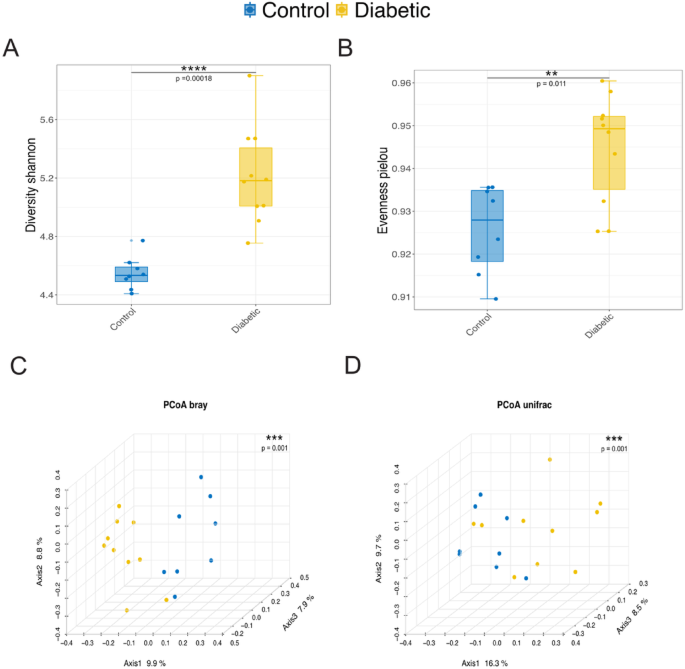
Microbial diversity and compositional differences between control and diabetic groups. (A) Boxplot illustrating the Shannon diversity index, reflecting overall microbial richness within the control and diabetic groups. The diabetic group exhibited significantly higher microbial diversity than the control group. (B) Boxplot of Pielou’s evenness, which measures the uniformity of species distribution. A higher evenness is observed in the diabetic group, indicating a more homogeneous distribution of microbial taxa relative to the control group. (C) Principal Coordinate Analysis (PCoA) plot based on Bray-Curtis dissimilarity, displaying the distinct clustering of microbial communities between control and diabetic groups along Axis 1, which explains 9.9% of the total variation. (D) PCoA plot based on unweighted UniFrac distances, further highlighting the separation of microbial communities between control and diabetic samples. Axis 1 accounts for 16.3% of the variation, demonstrating significant differences in phylogenetic composition. Alpha and beta diversity were assessed using the Wilcoxon rank-sum test and PERMANOVA, respectively.
Distinct shifts in gut microbial composition and abundance at family and fenus levels in diabetic versus control groups
To explore microbial diversity, a comprehensive taxonomic analysis was performed focusing on differences at both the family and genus taxonomic levels within the microbiome. The study revealed significant differences in the relative abundance of several microbial families between the diabetic and control groups (Fig. 2A). Notably, the diabetic group had a higher relative abundance of Bacteroidaceae and Lachnospiraceae, whereas Streptococcaceae was more abundant in the control group. These findings highlight distinct group-associated microbial patterns rather than temporal changes between the two groups.
Further differences were observed at the genus level (Fig. 2B). The genera Bacteroides, Blautia, and Streptococcus were significantly more abundant in the diabetic group. Conversely, genera linked to Lachnospiraceae and Ruminococcaceae, such as Faecalibacterium and Roseburia, were more prevalent in the control group. These distinctions at the genus level further emphasize the divergent microbial profiles between the two conditions. Additionally, other notable genera, such as Bifidobacterium, Methanobrevibacter, and Christensenellaceae, were distributed differently between the two groups. The diabetic group showed a higher relative abundance of Bifidobacterium, while Christensenellaceae was predominantly found in the control group.
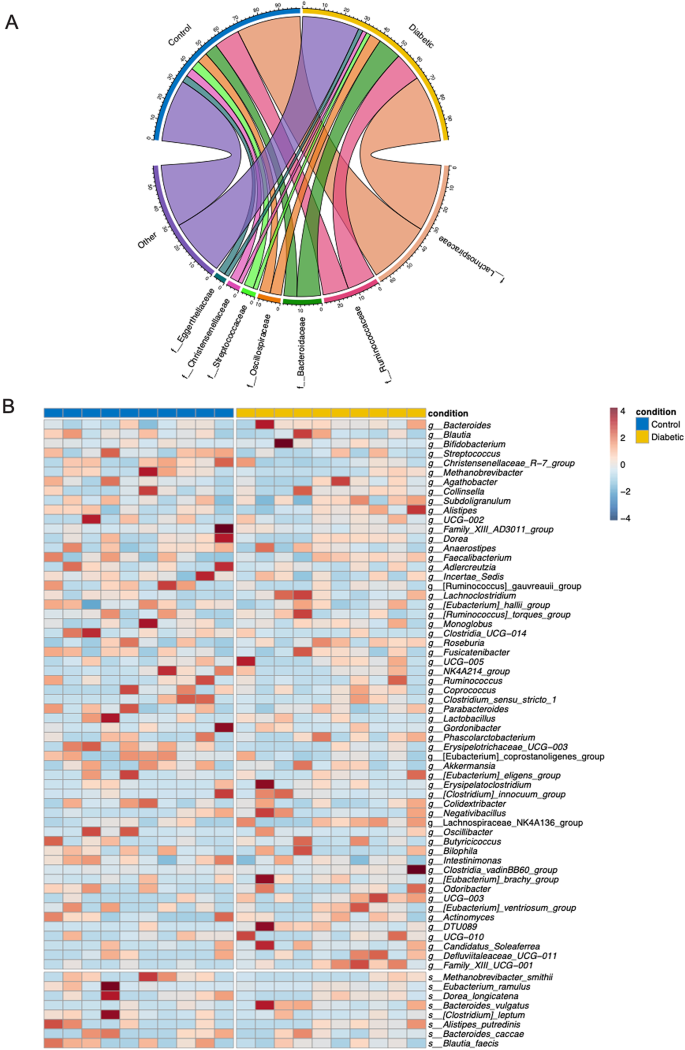
Differential microbial composition and abundance between control and diabetic groups. (A) Circos plot illustrating the differential microbial family-level composition between control and diabetic groups. Each segment represents a microbial family, with the connections between the groups showing the relative abundance of each family in the two conditions. Diabetic patients show a higher abundance of families such as Bacteroidaceae and Lachnospiraceae, while families such as Streptococcaceae are more prevalent in the control group. (B) Heatmap showing the genus-level relative abundance of bacterial taxa across control (blue) and diabetic (yellow) groups. The intensity of the colors represents the relative abundance of each genus, with red indicating higher abundance and blue indicating lower abundance. Notably, genera such as Bacteroides, Blautia, and Streptococcus are more abundant in diabetic patients, whereas Lachnospiraceae and Ruminococcaceae show higher abundance in the control group. This analysis highlights significant differences assessed by Wilcoxon test and p-value were corrected using FDR (p < 0.05) in the microbial composition between the two conditions, suggesting a potential role of specific bacterial taxa in the pathophysiology of diabetes.
LEfSe highlights core microbial taxa underpinning gut microbiome divergence in diabetic and control groups
LEfSe analysis identified significant differences in bacterial taxa between the control and diabetic groups. Taxa from the families Bacteroidaceae and Erysipelotrichaceae were predominantly enriched in the diabetic group. In contrast, Streptococcaceae was more abundant in the control group, as illustrated by the phylogenetic cladogram (Fig. 3A). Supporting these observations, the LDA scores further delineated the microbial differences between the groups. Taxa with positive LDA scores, including Bacteroides and members of the Lachnospiraceae family, were predominantly associated with the diabetic group. Conversely, negative LDA scores pointed to taxa like Streptococcus and Leuconostoc, which were more abundant in the control group (Fig. 3B). Additionally, boxplots displaying the relative abundance of key bacterial genera provided a clear illustration of the divergences between the groups (Fig. 3C). Genera such as Bacteroides, Lachnospiraceae_FCS020_group, and Angelakisella were significantly enriched in the diabetic group, suggesting their potential involvement in disease-related microbial shifts. On the other hand, the genera Leuconostoc and Streptococcus were more prevalent in the control group, reinforcing the distinct microbial profiles of healthy individuals. The statistical significance of these findings further highlights the robustness of the observed differences in bacterial communities between the diabetic and control groups.
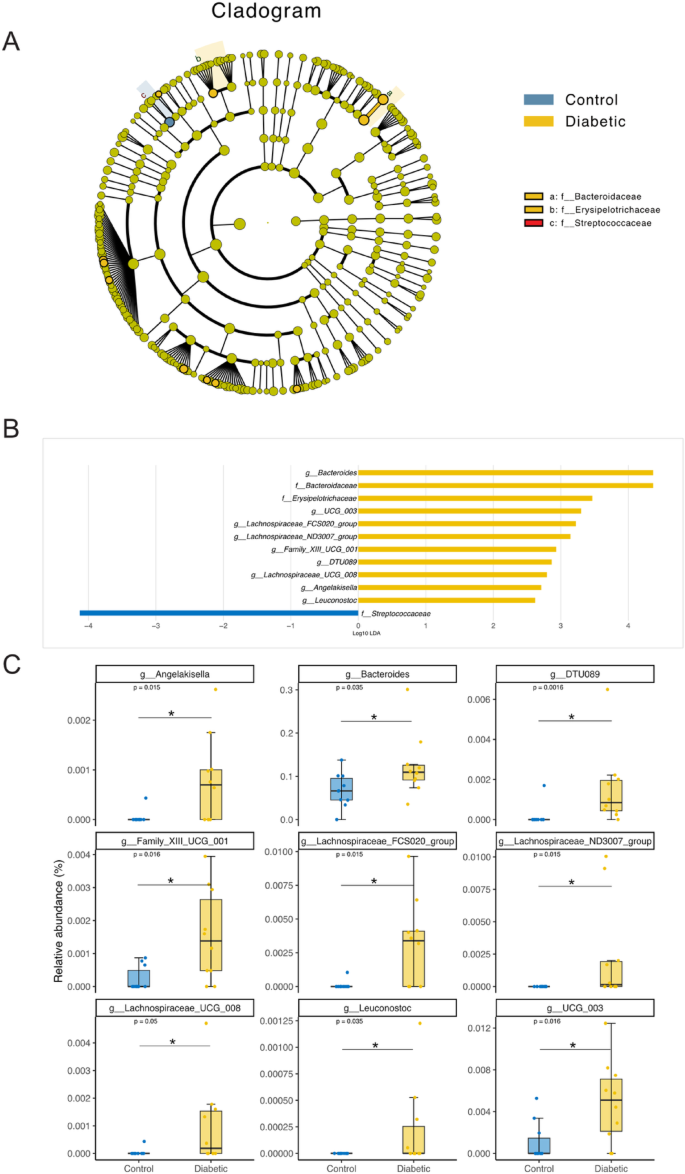
LEfSe analysis of bacterial taxa distinguishing control and diabetic groups.
(A) Cladogram showing the phylogenetic distribution of bacterial taxa enriched in control (blue) and diabetic (yellow) groups based on Linear Discriminant Analysis Effect Size (LEfSe). Taxa enriched in the diabetic group are primarily from the families Bacteroidaceae and Erysipelotrichaceae, while Streptococcaceae is more abundant in the control group. (B) Linear Discriminant Analysis (LDA) scores highlighting bacterial taxa with significant differential abundance between the control and diabetic groups. Positive LDA scores (yellow bars) represent taxa enriched in the diabetic group, while negative LDA scores (blue bars) represent taxa enriched in the control group. (C) Boxplots of the relative abundance of key bacterial genera identified by LEfSe analysis. Genera such as Bacteroides, Lachnospiraceae_FCS020_group, and Angelakisella are significantly more abundant in diabetic patients, while Leuconostoc and Streptococcus are more prevalent in the control group. Statistical significance is indicated with adjusted p-values (p < 0.05) using Wilcoxon test for each comparison.
Distinct microbial pathways and taxa associations characterize metabolic alterations in diabetic gut microbiomes
Of the 375 predicted metabolic pathways, 233 showed significant differences between the control and diabetic groups. To explore the association of these pathways with specific bacterial genera altered in the diabetic group, participants were initially divided into control and diabetic groups. A correlation analysis was subsequently conducted. A strict threshold was implemented to ensure robust and meaningful associations: only pathways with a p-value below 0.0001 were included in the final correlation analysis. This process resulted in 159 pathways correlated with 59 altered bacterial genera. The analysis was further refined by applying a correlation cutoff of 0.7, along with a p-value threshold of less than 0.005. This aided in identifying the strongest associations between metabolic pathways and bacterial genera. These results demonstrated the differences between bacterial genera and metabolic pathways under the two conditions. While Bacteroides and Alistipes were positively correlated with seven different pathways in the control group, Fusicatenibacter was negatively correlated with three distinct pathways (Fig. 4A). Conversely, in the diabetic group, we found that Christensenellaceae, Eubacterium coprostanoligenes group, and UCG-005 positively contributed to most of the altered pathways (Fig. 4B).
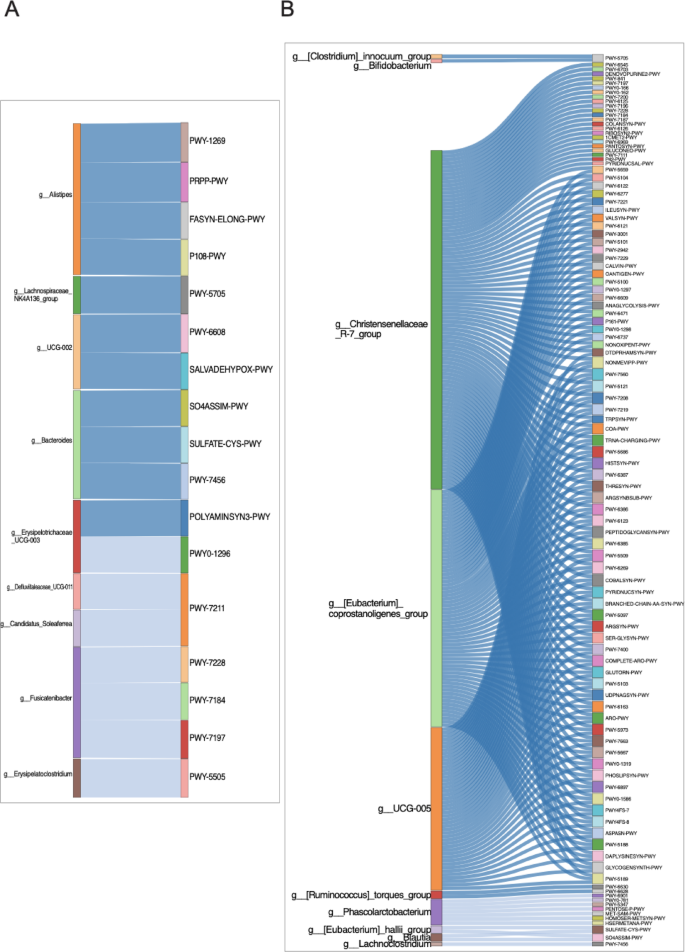
Functional pathway predictions and microbial taxa associations in control and diabetic groups. (A) Control group. (B) Diabetic group. Each Sankey plot illustrates significant correlations (adjusted p < 0.05) between differentially abundant microbial genera and functionally enriched metabolic pathways. Correlations were calculated using Spearman’s method, with a threshold of |r| ≥ 0.7. Dark blue ribbons indicate strong positive correlations (r > 0.7), while light blue ribbons represent strong negative correlations (r < − 0.7). Only taxa and pathways with both statistical significance and functional relevance are displayed, highlighting distinct microbe–function associations in each group.
Metabolomic profiles in diabetic versus control groups
Metabolomic analysis revealed differences in key metabolic pathways between diabetic and control groups. Pathway enrichment analysis identified fatty acid metabolism, glucose regulation, bile acid metabolism, and amino acid biosynthesis as significantly enriched in the diabetic group (Fig. 5A). Pathways related to fatty acid metabolism, particularly fatty acid elongation and β-oxidation, were more pronounced in diabetic individuals. These pathways displayed a high enrichment ratio, indicating changes in lipid metabolism in the diabetic group compared to controls (Fig. 5B). Additionally, pathways related to glucose homeostasis, such as insulin signaling and hexose metabolism, were enriched in the diabetic group, reflecting differences in glucose metabolic processes. Bile acid metabolism pathways were notably enriched in the diabetic group. Specifically, primary bile acid synthesis showed a significant enrichment ratio, pointing to potential changes in bile acid processing in diabetic individuals. Amino acid biosynthesis and transport pathways, including those for branched-chain amino acids (BCAAs) and aromatic amino acids, were also significantly enriched, suggesting alterations in amino acid metabolism in the diabetic group. The heatmap of metabolite abundances (Fig. 5C) revealed distinct clustering between the diabetic and control groups. Diabetic individuals displayed higher metabolites associated with the enriched pathways, including metabolites linked to fatty acid metabolism, glucose regulation, and bile acid synthesis. This clustering pattern supports the significant differences in metabolite profiles between the groups.
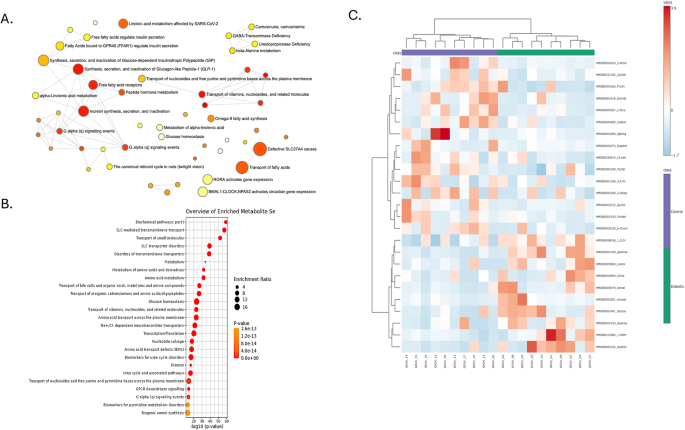
Metabolite Pathway Enrichment and Abundance Differences between Diabetic and Control Groups. (A) Network plot illustrating enriched metabolic pathways in diabetic versus control groups. Nodes represent pathways, with node size corresponding to the enrichment ratio and node color indicating significance (p-value). Pathways related to fatty acid metabolism, glucose homeostasis, and amino acid biosynthesis significantly enrich the diabetic group. (B) Bar plot showing the top enriched metabolic pathways based on enrichment ratios and p-values. Pathways such as bile acid metabolism, glucose homeostasis, and amino acid transport are highly enriched in diabetic samples. (C) Heatmap representing the relative abundance of key metabolites across control and diabetic groups. Red indicates higher abundance, and blue represents lower abundance. Clustering shows distinct metabolite profiles between control and diabetic individuals, highlighting potential biomarkers and metabolic shifts associated with diabetes.
Co-abundance correlation analysis reveals restructured microbial interactions in diabetes
To elucidate potential alterations in microbial ecological structure associated with type 2 diabetes, we performed a co-abundance correlation analysis based on Spearman’s rank correlations with FDR-adjusted p-values. This analysis revealed a substantial shift in microbial interaction dynamics, with the most significant correlations being positive (73.1%) and a smaller fraction negative (26.9%) (Fig. 6C). The predominance of positive correlations suggests enhanced co-occurrence and potential cooperative behavior among microbial taxa in the diabetic gut environment. Taxa belonging to the phyla Firmicutes and Bacteroidota were among the most interconnected, exhibiting the highest number of statistically significant associations (Fig. 6A). Furthermore, degree distribution analysis indicated that several taxa serve as highly connected nodes, or potential hubs, within the microbial network (Fig. 6B), implying their key role in structuring microbial communities in diabetes. These findings indicate that diabetes is not only characterized by taxonomic and functional shifts but also by a reorganization of microbial interaction networks – hallmarks of ecological dysbiosis that may influence host metabolic outcomes.
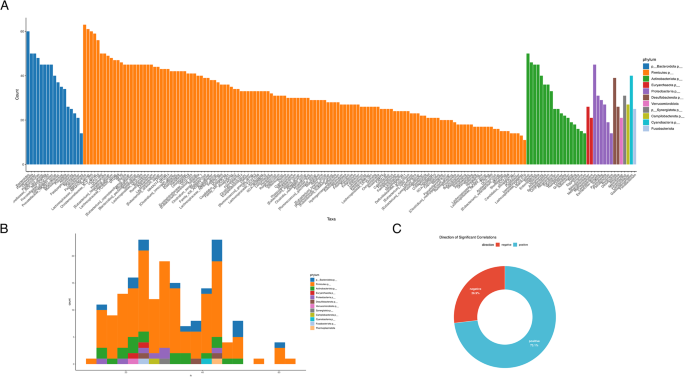
Co-abundance correlation analysis of microbial taxa in diabetic subjects. (A) Bar plot showing the number of statistically significant pairwise correlations per taxon, colored by phylum. The height of each bar represents the number of significant associations that the taxon forms with others (Spearman correlation, FDR-adjusted p < 0.05). (B) Histogram depicting the distribution of taxa by degree (number of connections), stratified by phylum. (C) Donut chart illustrating the directionality of significant correlations. Positive correlations (blue) accounted for 73.1%, while negative correlations (red) represented 26.9%, indicating that most microbial interactions in the diabetic group were co-occurring rather than mutually exclusive.
link