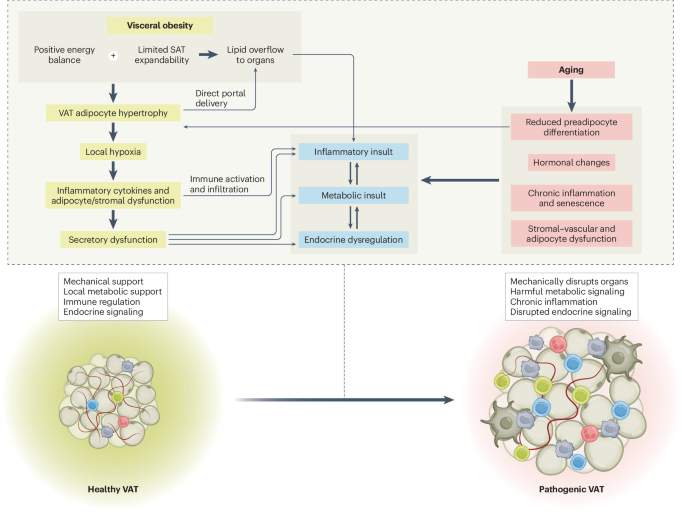
AQP7 deficiency drives adipose tissue remodeling and disrupts homeostasis
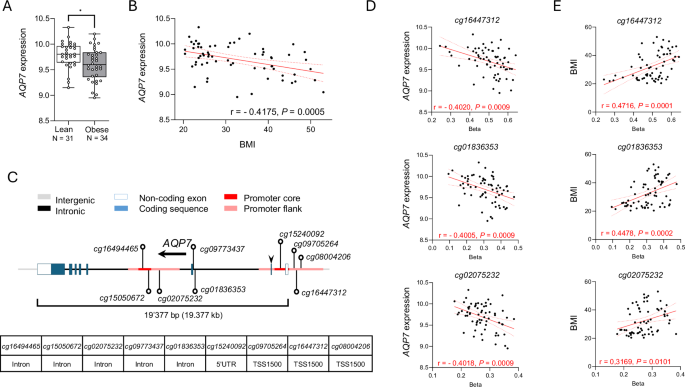
CpG methylation in human adipose tissue
CpG methylation and microarray gene expression data for the AQP7 locus were extracted from a previously published cohort examining the global CpG methylation profile in subcutaneous white adipose tissue of women spanning a range of BMI55. From this cohort, 65 women were included where AQP7 CpG methylation and gene expression were available. Briefly, an abdominal subcutaneous white adipose tissue biopsy was obtained by needle aspiration at 8:00 AM after an overnight fast. Gene expression was performed as described on whole white adipose tissue pieces using the Gene 1.0 or 1.1 ST Affymetrix arrays (Affymetrix, Inc., Santa Clara, CA)56. For global CpG methylation, the adipocytes were first isolated using collagenase digestion before DNA isolation. The Infinium Human Methylation EPIC BeadChip (Illumina, San Diego, CA) was used for CpG profiling55. Obesity was defined as a BMI > 30, while a lean/healthy BMI is considered < 25, according to the WHO57. The study was approved by the regional ethics board in Stockholm, and written informed consent was obtained from each subject.
Experimental animals and a high glucose diet
Knock-out (Aqp7−/−), haplodeficient (Aqp7+/−) and wild-type (Aqp7+/+) littermates (C57BL/6J background; aged 6 to 8 weeks)10,58 were used to analyse the effect of varying AQP7 expression in a glucose-induced obesity model59. The mice were randomized into two groups: one group received water supplemented with 30% (vol/vol) dextrose, while the other group received plain water without any supplementation for 8 weeks (Supplementary Fig. 1A). All the mice were allowed free access to a standard laboratory diet (Scientific animal food & engineering, Augy, France). Food and water intake, as well as body weight, were measured once per week. All mice were housed in an air-conditioned room with a 12/12-h dark-light cycle. The experiments were conducted in accordance with the National Research Council Guide for the Care and Use of Laboratory Animals and approved by the ethics committee of the UCLouvain Medical School (Brussels, Belgium).
Micro-computed tomography analysis
Micro-CT scans were performed on mice anesthetized with isoflurane/oxygen to monitor changes in body composition and fat mass. Scanning was performed with Skyscan 1278 (Bruker, Kontich, Belgium) at 50 µm voxel resolution using a source voltage of 65 kV and a current of 770 uA as described60. Raw images were then reconstructed with an isotropic voxel size of (51 × 51 × 51) µm³.
Whole-body composition analysis in mice was performed using micro-CT to quantify adipose tissue, lean tissue, and skeletal volumes. Visceral fat area was assessed at the L4 level via single-slice analysis, using semi-automatic region-of-interest (ROI) delineation and a grey value threshold of 30–57. Dorsal muscle analysis involved manual ROI selection at L4 and L5, applying a grey value threshold (30–97) to exclude non-lean tissue, with muscle density subsequently measured. Virtual liver biopsy was conducted by placing a 3D cylindrical ROI in the liver, and the mean density (HU) was automatically computed. Dorsal muscle and liver densities were normalized to spleen density as an internal control (Supplementary Fig. 1B). All analyses were performed using SkyScan software (CT Analyser version 1.17.7.2).
Peritoneal transport studies, blood and tissue sampling
A peritoneal equilibration test (PET) with 2.5 mL of 3.86% glucose dialysate was performed to evaluate peritoneal transport parameters, as previously described61. Briefly, the mice underwent a 2-h PET following the administration of 2.5 mL of 3.86% glucose dialysate. After the dwell, the dialysate was collected from the peritoneal cavity, and the net ultrafiltration (UF) volume was determined. Mice were euthanized using an overdose of sevoflurane, in accordance with approved ethical guidelines. Blood samples were collected from the inferior vena cava and immediately centrifuged to separate the serum. Serum was stored at −20 °C until analysis. Triglycerides and urea were assayed using Dri Chem NX500i (Fujifilm, Tokyo, Japan). The visceral peritoneum was snap-frozen in liquid nitrogen and stored at −80 °C, or routinely fixed in 4% paraformaldehyde in 0.1 M phosphate buffer (VWR, Brussels, Belgium). Protein concentration was assayed using the Pierce BCA protein assay kit (Thermo Fisher Scientific, Waltham, MA, USA).
MEFs cell culture and differentiation
Mouse embryonic fibroblasts (MEFs) were prepared from 13.5 days Aqp7+/+ and Aqp7−/− mouse embryos as previously described27. The embryos were extracted from the uterus and minced with a sterile scalpel under a cell culture hood. Single cell suspensions were prepared by incubating for 20 min at 37 °C with 0.25% trypsin/EDTA. The reaction was stopped with the addition of medium containing 10% fetal bovine serum (FBS). DNase was added at the final concentration of 100 μg/ml and incubated for 15 min at 37 °C. Cells were grown in 5% CO2 incubator at 37 °C in culture medium (high glucose DMEM) supplemented with 10% (v/v) FBS. Adipogenesis was induced in two days postconfluent cells using a well-established combination of proadipogenic factors: 1 μM dexamethasone, 0.5 mM 3-isobutyl-1-methylxanthine (IBMX), 5 μg/ml insulin, and 0.5 μM rosiglitazone in pre-differentiation medium (high glucose DMEM supplemented with 10% FBS). Cells were exposed to these factors for 2 days, following which the medium was substituted with post-differentiation medium (high glucose DMEM supplemented with 10% FBS) containing only 5 μg/ml insulin. Post-differentiation medium was renewed every other day until day 12. On day 12 post-differentiation, the supernatants from both Aqp7+/+ and Aqp7−/− cell cultures were collected and used for further analysis. Wild-type fibroblasts were subsequently exposed to the 0.22-μm-filtered supernatants from both Aqp7+/+ and Aqp7−/− cells for 12 h. After the exposure period, the cells were harvested and processed for downstream assays to evaluate the effects of the different conditioned media.
Lipid visualization/quantification using Oil Red staining
After being washed twice with PBS, the differentiated cells were fixed for 15 min at room temperature using 4% paraformaldehyde (PFA) in PBS. Subsequently, the fixed cells underwent two washes with dH2O, one wash with 60% isopropanol, and were then stained for 5 min with 0.2% Oil Red O in 60% isopropanol. Excess stain was removed by washing three times with tap H2O. Images were captured using an optical phase contrast microscope fitted with a digital camera. Spectrophotometric evaluation of the dye (Abs 500 nm) was carried out after lipid extraction using 100% isopropyl alcohol for 10 min.
Enzyme-linked immunosorbent assays
The levels of insulin (Mercodia, Uppsala, Sweden), MCP-1 (Sigma-Aldrich, St Louis, MO, USA), IL-6 and IL-1β (R&D Systems, Wiesbaden-Nordenstadt, Germany) in serum or dialysate samples were quantified in duplicate using Enzyme-linked immunosorbent assay, according to the manufacturer’s protocols.
Glycerol, free fatty acid, and lactate analysis
Colorimetric assay kits were used to quantify glycerol (#MAK117, Sigma-Aldrich), free fatty acid (#MAK044-1KT, Sigma-Aldrich), and lactate (#MAK064, Sigma-Aldrich) released from adipocytes in serum or cell culture medium samples and performed as per the manufacturer’s instructions.
RT-PCR and quantitative real-time RT-PCR
Total RNA from mouse visceral peritoneum was extracted with Trizol (Invitrogen, Merelbeke, Belgium) and using AurumTM Total RNA fatty and fibrous tissue kit (Bio-Rad, Hercules, CA, USA) according to the manufacturer’s protocol. DNase I treatment was performed to eliminate genomic DNA contamination. One microgram of RNA was used to perform the reverse-transcriptase reaction with iScriptTM cDNA Synthesis Kit (Bio-Rad). Three reference genes (Gapdh, Actb, and 36b4) not influenced by treatment protocols were used for the normalization. The sequences and efficiencies of the primers are given in (Supplementary Table 1). Changes in the mRNA levels of target genes were determined by semiquantitative reverse-transcriptase-polymerase chain reaction (RT-PCR) with an iCycler IQ system (Bio-Rad, Hercules, CA, USA) using SYBR Green I detection as described previously61. The PCR conditions were 95°C for 3 min followed by 40 cycles of 30 s at 95 °C, 15 s at 60 °C and 1 min at 72 °C. The relative changes in target mRNA between groups were determined by using the relation 2−ΔΔCt.
Tissue staining and immunofluorescence
Samples from the peritoneum were fixed in 4% paraformaldehyde (PAF), embedded in paraffin and cut into 5 µm sections. Hematoxylin-eosin (HE), picrosirius red staining and immunostaining were performed as previously described10. The areas of adipocytes in the visceral peritoneum (HE) were calculated (π∙radius²) in at least 50 cells per section, and the cells were automatically counted using ImageJ software. Sirius red staining was observed under polarized light microscopy (AXIOSCAN; Zeiss, Oberkochen, Germany). The area of collagen fibers as a function of their color was quantified from tissue sections stained62. The color corresponds to relative fiber thickness from thin green fibers to increasingly thick yellow, orange, and red fibers.
For immunofluorescence, the tissue sections were deparaffinized and rehydrated using a series of ethanol dilutions of decreasing concentration. Antigen retrieval was performed using sodium citrate buffer (10 mM, in distilled water, pH 6.0). The tissue sections were heated in the microwave at 900 W for 4 min, followed by 90 W for 15 min, then 900 W for 1 min 30 s and finally cooled at room temperature for 10 min. The slides were then blocked with blocking buffer (1% BSA, 0.2% non-fat dry milk in PBS) for 60 min and incubated with primary antibody in blocking buffer overnight at 4 °C. After three washes in 0.1% Tween 20 (v/v in PBS), the tissue slides were incubated with fluorophore-conjugated secondary antibody (Life Technologies, Carlsbad, CA, USA) diluted in blocking buffer at room temperature for 1 h. The complete list of antibodies used for immunofluorescence is included in Supplementary Table 2. Finally, the slides were mounted in Prolong Gold anti-fade reagent (Invitrogen, Waltham, MA, USA) and examined using a Zeiss LSM 510 Meta Confocal microscope (Carl Zeiss, Jena, Germany).
Proteomics analysis of cell culture supernatants (secretome analysis)
Sample preparation
Following experimental treatments, supernatants were collected and frozen until analysis at −80 °C. Combinatorial peptide ligand library (CPLL) equalizer beads (ProteoMiner, Bio-Rad, Hercules, CA, USA) were prepared per manufacturer’s protocol. About 3 mL supernatant were mixed with 600 μl CPLL bead solution (150 μl bead bed volume) and incubated on a roller mixer (overnight, 4 °C). Total protein concentration was determined (Pierce 660 nm Protein Assay, Thermo Fisher Scientific) per the manufacturer’s manual. Pure FBS samples were processed in the same way as the samples in order to generate an exclusion list for bovine proteins contained in the supernatant.
Protein digestion and labelling
About 110 µg of each sample were used. Before digestion, the pH was adjusted to neutral. Digestion was performed using single-pot, solid-phase enhanced sample preparation (SP3). Briefly, the reduced (10 mM DTT for 1 h at 56 °C) and alkylated (55 mM IAA, 30 min at RT) proteins were bound to SP3 beads (10:1 beads:protein ratio, GE Healthcare, Illinois, USA), washed with 80% ethanol and acetonitrile, and subjected to on-bead digestion with trypsin/LysC (1:25 protease:protein ratio, Promega, Madison, Wisconsin, USA) overnight at 37 °C in 50 mM ammonium bicarbonate, pH 8.5 (Sigma-Aldrich, Schnelldorf, Germany). Following elution and desalting (Peptide Desalting columns; Thermo Fisher Scientific), peptides were dried in a vacuum concentrator. Peptide concentration was determined (Colorimetric Peptide Assay, Thermo Fisher Scientific) according to the manufacturer’s protocol. An internal pooled standard was prepared of equal parts of all samples.
Peptides were labeled with isobaric mass tags for multiplexing (TMTpro, Thermo Fisher Scientific) according to the instructions provided by the manufacturer, with minor modifications. TMTpro reagents were reconstituted with acetonitrile and 70 µg each sample were labeled with 210 µg of TMTpro reagent. After incubation for 1 h at RT the reaction was quenched by the addition of 5% hydroxylamine (Sigma-Aldrich) in TEAB for 15 min at RT. Labeling efficiency was determined by LC-MS.
Offline fractionation
Pooled labeled samples were concentrated and desalted (Pierce Peptide Desalting Columns; Thermo Fisher Scientific). Eluates were dried in a vacuum concentrator and reconstituted (20 mM ammonia formate, pH 10) prior to fractionation at basic pH. All fractionations were performed using an UltiMate 3000 system (Thermo Fisher Scientific). Peptides were separated on a Gemini-NX C18 (150 × 2 mm, 3 µm, 110 A, Phenomenex, Torrance, USA) column in 20 mM ammonia formate buffer, pH 10.
Liquid chromatography–mass spectrometry (LC-MS)
Fractions were analyzed on an Ultimate 3000 RSLC nano coupled directly to an Exploris 480 with FAIMSpro (all Thermo Fisher Scientific). Samples were injected onto a reversed-phase C18 column (50 cm×75 µm i.d., packed in-house) and eluted with a gradient of 4 to 38% mobile phase B over 94 min by applying a flow rate of 230 nl/min. MS scans were performed in the range from m/z 375–1650 at a resolution of 120,000 (at m/z = 200). MS/MS scans were performed, choosing a resolution of 30,000 with the turboTMT mode for TMpro Reagent; normalized collision energy of 33%; isolation width of 0.7 m/z and dynamic exclusion of 90 s. Two different FAIMS voltages were applied (−40 and −60 V) with a cycle time of 1.5 s per voltage. FAIMS was operated in standard resolution mode with a static carrier gas flow of 4.6 L/min.
Spectral analysis
The acquired raw MS data files were processed and analyzed using ProteomeDiscoverer (v2.4.0.305, Thermo Fisher). SequestHT was used as a search engine, and the following parameters were chosen: database: Mus musculus (SwissProt, downloaded on 2022-06-15) as well as the bovine fasta file created from the analysis of FCS; enzyme: trypsin; max. missed cleavage sites: 2; static modifications: TMTpro (K and peptide N-terminus) and carbamidomethyl (C); dynamic modifications: oxidation (M), acetyl (protein N-terminus), Met-loss (M) and Met-loss + acetyl (M); precursor mass tolerance: 10 ppm; fragment mass tolerance: 0.02 Da. For reporter ion quantification, the most intense m/z in a 20 ppm window around the theoretical m/z was used. Correction of isotopic impurities for reporter ion intensities was applied. Only unique peptides, above an average S/N threshold of 10, were used for quantification. Normalization was based on total peptide amount and scaling mode on the control average (internal standard) in the first round. All Master Proteins and Master Protein candidates with FDR <0.01 were exported, and the assignment to a species was checked. A fasta file was created out of all mouse proteins with more than one matching peptide and used in a second analysis for normalization of the data. Afterwards, all Master Proteins and Master Protein candidates with FDR <0.01 were exported, and the assignment to a species was checked. Single peptide IDs were excluded from the dataset, and only peptides and proteins with FDR <0.01 were reported.
Data analysis
Results are expressed as mean ± SD. Differences between groups were assessed by unpaired t-test, one-way ANOVA followed by Tukey’s multiple comparisons test or two-way ANOVA (to assess genotype effect in different time points), using GraphPad Prism Software version 9.4.0 (San Diego, California, USA). A p value <0.05 was considered statistically significant.
Statistical analyses of the proteomics data and graphical representations of results were performed using R (v4.0.3; Ingenuity Pathway Analysis (IPA 7.0, Qiagen, or Reactome Pathway Analysis ( were used to identify involvement of pathways, their respective predicted activation or deactivation patterns, and their enrichment based on differentially abundant proteins (one-tailed Fisher’s exact test, alpha 0.05). The IPA-calculated z-score assessed the activation state based on observed and predicted up/downregulation patterns, while Reactome pathway analysis provided insights into the biological relevance of the dataset by mapping proteins to known curated pathways. Differential protein abundances were analyzed with linear models for microarray data (LIMMA) using the R package “limma” as described before63. The Human Protein Atlas64 was used to extract cell-specific proteins of the most common cell types in adipose tissue. These proteins were then filtered for receptors according to Browaeys et al40. For cell-specific patterns of communications, receptors present in all four cell types were removed from the analysis.
link