Advancing sleep health equity through deep learning on large-scale nocturnal respiratory signals
Datasets and model training
In our study, four public datasets from the National Sleep Research Resource (NSRR)34, namely the Sleep Heart Health Study (SHHS)35, the Multi-Ethnic Study of Atherosclerosis (MESA)36, the MrOS Sleep Study (MrOS)37, and the Study of Osteoporotic Fractures (SOF)38 were employed. Table 1 provides a summary of these datasets, including the number of nights, age groups, male proportions, AHI distribution, and disease conditions. Detailed descriptions of each dataset are presented in the Supplementary Notes. Although these public datasets encompass individuals with varying degrees of SDB severity and both sexes, they underrepresent individuals under 45. Meanwhile, recent studies have revealed a growing prevalence of SDB among younger and middle-aged populations (18–45)39,40,41,42, with strong associations with cardiovascular and metabolic comorbidities43,44. From a technical perspective, this age imbalance may limit model generalization. To develop a universally applicable sleep assessment model, we prospectively collected a multi-center clinical dataset covering over 1000 nights from younger adults (under 45 years), including both respiratory belt-derived respiratory signals (ClinSuZhou and ClinHuaiAn) and radar-derived respiratory signals (ClinRadar).
We developed a deep-learning model named ResSleepNet for automatic sleep staging and AHI estimation. Figure 1 illustrates the model’s framework and the adversarial learning process on the internal dataset. Figure 2 offers a more detailed explanation of the model training and inference process used in this study. Phase I encompasses the pre-training process, during which we train the model on the internal dataset by minimizing the losses for sleep staging and AHI estimation while maximizing the loss for the domain discriminator. In phase II, the trained model weights are frozen, and performance evaluation is carried out on the external dataset. For details on data pre-processing and model training, validation, and testing, refer to the Methods section.
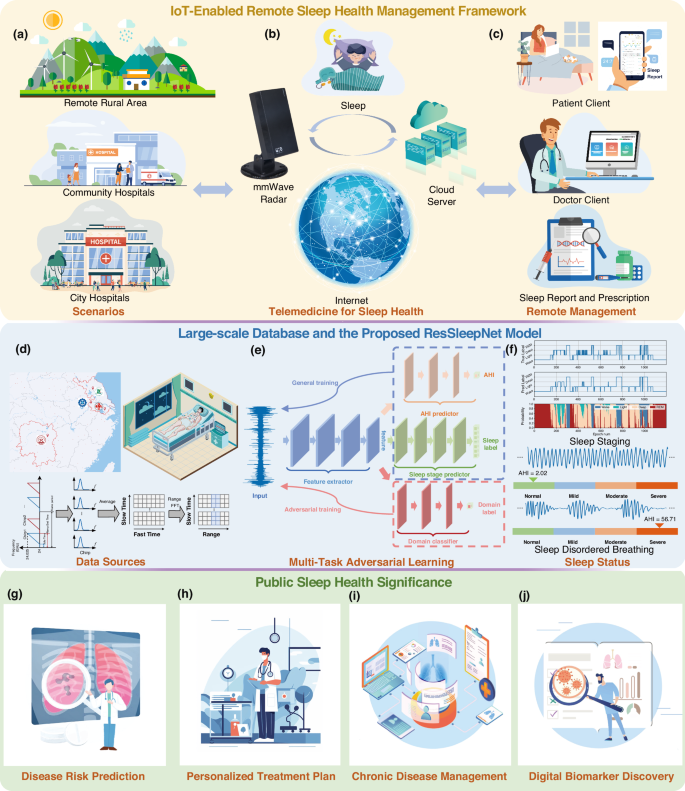
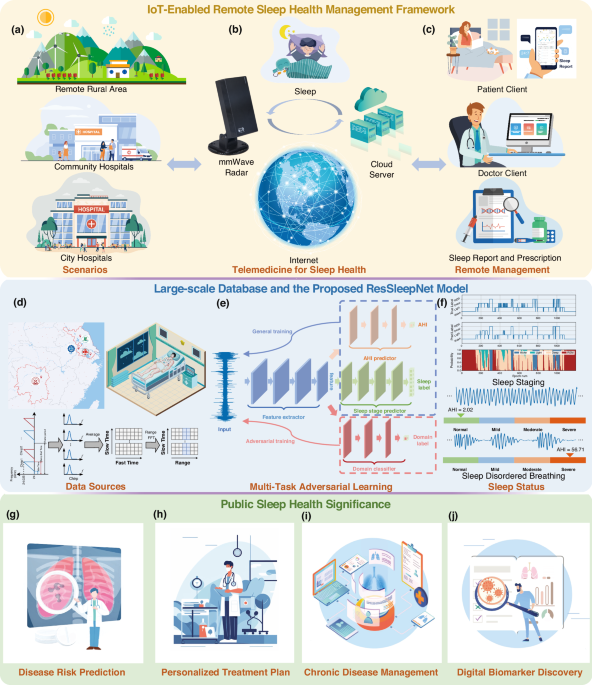
a Deployment scenarios, including remote rural areas, community hospitals, and city hospitals. b Data acquisition using mmWave radar and data transmission to the cloud server. c User interfaces for patients and doctors to access sleep reports and manage treatments. Large-scale Datasets and the Proposed ResSleepNet Model: d Data sources used for model development. e Multi-task adversarial learning framework for sleep staging and AHI prediction. f Sleep status assessment results, including sleep staging and analysis of sleep-disordered breathing. Public Sleep Health Significance: g predicting disease risks associated with sleep disorders, h formulating personalized treatment plans tailored to individual patient needs, i managing chronic diseases influenced by sleep problems, and (j) discovering novel digital biomarkers to support evidence-based medicine and precision healthcare.
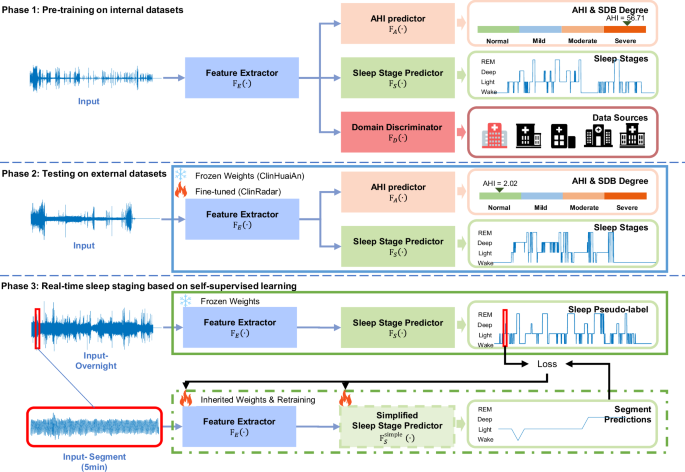
In Phase 1, the model is pre-trained on internal datasets using thoracoabdominal motion signals recorded from respiratory belts. The feature extractor FE( ⋅ ) first processes the input sleep signals, which extracts relevant features. These features are then fed into two predictors: the AHI predictor FA( ⋅ ) to assess the severity of sleep apnea, and the Sleep stage predictor FS( ⋅ ) to predict different sleep stages. A domain discriminator FD( ⋅ ) is also employed to distinguish data sources, enhancing the model’s generalization capability. In Phase 2, the pre-trained network (including the feature extractor, the AHI predictor, and the Sleep stage predictor) is applied with frozen weights to respiratory belt-derived respiratory signals (ClinHuaiAn) and fine-tuned for radar-derived respiratory signals (ClinRadar). In Phase 3, the model is employed for real-time sleep staging. The input signals are divided into overnight signals and 5-minute segments. The overnight signals are processed by the feature extractor and the sleep stage predictor with frozen weights to generate pseudo-labels. These pseudo-labels, together with the segment signals, are used for further fine-tuning of the feature extractor. A simplified sleep stage predictor is then used for real-time sleep stage prediction.
Sleep staging and AHI prediction
Table 2 presents the model’s performance across different datasets, including overall accuracy in sleep staging, the sensitivity of predictions for each sleep stage, and the accuracy of AHI estimation. In the internal test datasets, the sleep staging task attained the highest average accuracy of 83.53% and Kappa of 0.73 in MrOS. Performance was somewhat attenuated yet remained acceptable in SHHS (80.34%, 0.70), MESA (83.09%, 0.73), SOF (78.57%, 0.68), and ClinSuZhou (81.72%, 0.71). In the external datasets, the ClinHuaiAn dataset achieved an average accuracy of 79.62% and Kappa of 0.67, while the radar dataset showed a decrease with an accuracy of 75.81% and Kappa of 0.62. Owing to differences in sensing methods, radar-based thoracoabdominal motion signals can slightly differ from respiratory belts, particularly during sleep posture changes. Although the frequency remains consistent, signal amplitude may vary, resulting in reduced performance when transferring a respiratory belt-based pre-trained model to radar data. Nevertheless, radar monitoring in this study still achieves comparable performance.
We systematically compared the correspondence between the sleep stages predicted by ResSleepNet and the actual sleep stages. Figure 3 and Supplementary Fig. 1 display the confusion matrices of sleep staging for each dataset, while Supplementary Table 1 further analyzes the sensitivity and specificity (as defined in Supplementary Table 2) distribution of predictions for different sleep stages across datasets. In the internal datasets, ResSleepNet performed satisfactorily in recognizing wake, REM, and light sleep. However, its accuracy in detecting deep sleep was relatively lower, with frequent misclassifications of light sleep. The reasons are: (1) as shown in Supplementary Table 3, deep sleep makes up less than 15% of natural sleep, significantly lower than light sleep; (2) the similarity in cardiopulmonary features between deep and light sleep stages might further increase the difficulty of classification. To enhance the interpretability of the model, we visualized the intermediate decision-making process of the sleep staging prediction model in Supplementary Fig. 4. The results indicate that the model’s focus differs noticeably across wake, REM, and light/deep stages, though confusion tends to occur between light and deep stages, consistent with the analysis presented earlier. Additionally, the sensitivity for detecting deep sleep varies among different datasets. For instance, it reaches 68.97% in SHHS but only 46.09% in MrOS. As shown in Supplementary Table 3, the proportion of deep sleep throughout the night is a crucial factor influencing detection performance. In MrOS, the proportion of deep sleep is merely 6.14%, significantly lower than the 11.93% in SHHS. Other datasets also exhibit this trend, where the detection performance for deep sleep correlates with its proportion in total sleep time. However, in ClinRadar, the sensitivity for the wake and REM stages decreased significantly. The confusion matrix in Fig. 3(i) reveals that 21.26% of wake and 28.23% of REM stages were misclassified as Light sleep, which is higher than in the internal test sets and ClinHuaiAn. Despite the ClinRadar dataset comprising over 200 nights of data, it remains relatively small compared to other datasets, contributing to the overall decline in accuracy.
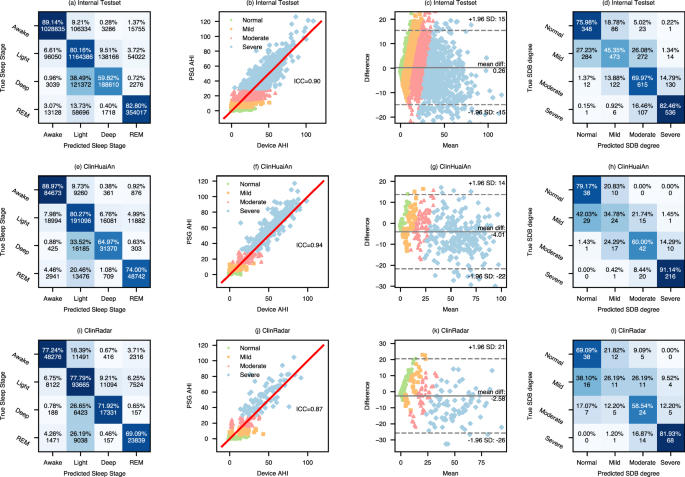
a, e, i Sleep stage confusion matrix, where the percentages indicate the proportion of correctly and incorrectly classified instances for each stage, and the numbers represent the actual counts of these classifications. b, f, j AHI scatter plot with the middle line representing the equation y = x. c, g, k Bland-Altman plot for AHI with horizontal lines for the mean difference and the 95% limits of agreement. d, h, l SDB severity confusion matrix. Source data are provided as a Source Data file.
In the AHI estimation task, the overall ICC of the internal test set was 0.90. With the exception of the SOF dataset, all other internal subsets achieved an ICC of 0.90 or higher, with ClinSuZhou attaining the highest value of 0.92. For the external datasets, the ICC values were 0.94 and 0.87, respectively. The middle two columns of Fig. 3 exhibit scatter plots and Bland-Altman plots that compare the true and predicted AHI values, demonstrating high consistency between the estimated and annotated AHI values. Supplementary Table 4 further analyzes other metrics used to evaluate AHI prediction performance and the results across different internal subsets. According to American Academy of Sleep Medicine (AASM) standards, the severity of SDB is classified based on AHI values into normal (AHI < 5), mild (5 ≤ AHI < 15), moderate (15 ≤ AHI < 30), and severe (AHI≥30). The four-class classification accuracies were 65.08% for the internal test set, 75.47% for ClinHuaiAn, and 63.80% for ClinRadar. The rightmost column of Fig. 3 presents the confusion matrix for the four severity categories. The results indicate that our model performs optimally in detecting severe SDB, achieving an accuracy of 91.14% in the ClinHuaiAn dataset. However, performance declines when identifying individuals with normal or moderate severity. The accuracy for detecting individuals with mild severity is the lowest, falling below 50% and dropping to under 40% in external datasets (as shown in the rightmost column of Fig. 3). Since the SOF dataset mainly consists of individuals with mild SDB (as shown in Table 1), the ICC for AHI estimation in SOF is only 0.77. Furthermore, we evaluated the tolerance accuracy, which allows a predicted severity level to differ from the true classification by no more than one category. In the internal dataset, 98.12% of patients were either correctly classified or off by just one severity level. The tolerance accuracy was 99.29% for ClinHuaiAn and 92.31% for ClinRadar (Supplementary Table 4). In the 3030 nights of the internal test set, only 2 nights were misclassified between the normal and severe categories (Fig. 3d), while no such misclassifications occurred in the two external datasets (Fig. 3h, l). This single-breathing-pathway-based model maintains reliable SDB monitoring accuracy while significantly enhancing patient comfort during sleep assessment.
To contextualize our model’s performance, we compare it against representative state-of-the-art (SOTA) HSAT approaches. As shown in Supplementary Table 6, our framework achieves competitive performance, benefiting from a larger dataset scale, robust model design, and enhanced portability for real-world applications.
Ablation study
To evaluate the contribution of transfer learning, we compared the performance of a model trained directly on radar-derived respiratory signals with a model pretrained on large-scale thoracoabdominal motion signals and subsequently fine-tuned on radar data. As shown in Supplementary Table 5, pretraining led to a 14.2% absolute improvement in accuracy (from 61.6% to 75.8%) and a notable increase in Cohen’s Kappa (from 0.37 to 0.62) for sleep staging. For AHI estimation, the ICC improved from -0.13 to 0.87, while the MAE was reduced by more than half (from 22.97 to 8.80 events/hour).
These improvements arise because large-scale respiratory belt datasets provide richer and more diverse respiratory patterns, enabling the model to learn robust low- and mid-level temporal features that transfer effectively to radar signals. Without this pretraining, the radar-only model struggles to capture such variability due to the relatively small radar dataset size. This two-stage strategy mitigates overfitting to the smaller radar dataset and significantly enhances cross-modality generalization, particularly for challenging sleep stages such as Deep and REM, and the AHI estimation task.
Sleep parameters and clinical correlation analysis
Sleep parameters
We performed a difference analysis on key sleep parameters, including total sleep time (TST), sleep efficiency (SE), sleep onset latency (SOL), wake after sleep onset (WASO), as well as the proportions of light sleep, deep sleep, and REM sleep. Detailed definitions of these parameters are provided in Supplementary Table 2. Figure 4a–g shows the violin plots of the differences in sleep parameters calculated from the true and predicted sleep labels. Each point in these plots represents the monitoring result of one night. The text above each violin plot indicates the correlation and P-value between the true and predicted sleep parameters. For all sleep parameters, the P-values are less than 0.0001, indicating strong statistical significance. The actual and predicted values for TST, SE, SOL, WASO, and REM sleep duration are highly consistent, with the corresponding error density plots concentrated around zero. In contrast, there is a slight deviation in the duration of light and deep sleep, and the error density plots have a broader distribution. The larger range in the density plots suggests misclassification between light and deep sleep stages, which leads to reduced accuracy in parameter estimation.
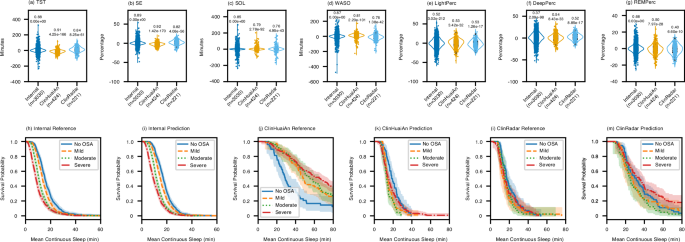
a–g Comparison of sleep parameters (TST, SE, SOL, WASO, and proportions of Light sleep, Deep sleep, and REM sleep) between the actual and predicted values across the internal dataset (n = 3030 nights), ClinHuaiAn dataset (n = 424 nights), and ClinRadar dataset (n = 221 nights). Each violin plot shows the distribution of the differences (prediction error) between the true and predicted values. The contour of each violin plot indicates the kernel density of these differences, while the horizontal line inside represents the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to the minimum and maximum values within 1.5 times the interquartile range (IQR). The numerical values above each plot represent the Pearson correlation coefficients between the actual and predicted parameters, and the significance of the correlations is assessed using a two-tailed Pearson correlation test with exact p-values. All n values refer to independent nights of sleep recordings. Source data are provided as a Source Data file. h–m Kaplan-Meier plots of the true and predicted average continuous sleep times in the internal dataset (h, i), the ClinHuaiAn dataset (j, k), and the ClinRadar dataset (l, m). The vertical axis indicates the survival probability (proportion of subjects with an average continuous sleep time greater than the duration shown on the horizontal axis). Different colored curves represent subjects in distinct sleep apnea-hypopnea severity categories (No OSA, Mild, Moderate, Severe). The shaded areas surrounding each curve denote the 95% confidence intervals of the survival probability estimates.
In addition to traditional sleep parameters, we also evaluated the model’s capability to assess sleep fragmentation. Figure 4h shows the Kaplan-Meier survival curve of the actual average continuous sleep duration in the internal dataset. Different colors represent specific SDB severity levels. Each curve indicates the proportion of subjects within a category who maintain continuous sleep beyond a certain duration. The results show significant differences in survival curves across SDB severity levels, with more severe SDB leading to increased sleep fragmentation. The predicted results in Fig. 4i closely follow this trend and exhibit good consistency with the actual results in Fig. 4h. However, due to the limited data volume, the Kaplan-Meier survival curves for actual average continuous sleep duration in the two external datasets show some deviation compared to those in the internal test set.
General clinical correlation analysis
To further illustrate the adaptability of our method to different groups, we analyzed the correlation between sleep staging accuracy and various general clinical information, including sex, age, body habitus (BMI), and the severity of sleep apnea.
Sex
We analyzed the relationship between single-night sleep staging performance and sex. As shown in Supplementary Fig. 5a, d, in the internal test set, which has the largest sample size and a relatively balanced male-to-female ratio, the model’s performance for males and females shows no significant difference, with similar performance distributions. In ClinHuaiAn (86.8% male) and ClinRadar (82.0% male), the performance distribution between males and females remains close, with slight differences attributable to the sex imbalance. Overall, the results indicate that the model’s performance does not differ between males and females.
Age
As shown in Table 1, we divided the subjects into four age groups: under 45, 45–55, 55–65, and over 65. Supplementary Fig. 6a, d show that in the internal test set, the accuracy and Kappa values are higher and more stable for participants under 45. As age increases, the median accuracy and Kappa gradually decrease, and the range of their distribution widens, indicating greater variability among older participants. A similar but more pronounced trend is observed in the external datasets (Supplementary Fig. 6b, e, c, f), with larger fluctuations, particularly in the older age groups (above 55). This suggests that the external datasets introduce additional variability, possibly due to differences in population characteristics or data collection environments. Overall, the model’s performance declines in older participants, likely due to changes in sleep architecture and the presence of comorbidities in older individuals, making accurate staging more challenging.
Body habitus
To assess the model’s robustness against inter-individual differences in body composition, we stratified participants by body mass index (BMI) into four conventional categories: underweight (UW, BMI < 18.5), normal weight (NW, 18.5≤BMI < 25), overweight (OW, 25≤BMI < 30), and obese (OB, BMI≥30). As shown in Supplementary Fig. 7 and Supplementary Fig. 8, the model maintained stable performance across all BMI categories and both sexes. Notably, despite potential concerns that excess adipose tissue in individuals with obesity could attenuate thoracoabdominal motion signals–especially in radar-based sensing–the staging accuracy in the obese group did not show significant degradation. These findings suggest that our preprocessing strategies (e.g., signal normalization) and model design contribute to mitigating variations introduced by body habitus, supporting reliable performance even in individuals with elevated BMI.
Sleep Apnea
We categorized the subjects into four groups based on their clinical AHI values: normal, mild, moderate, and severe SDB (as shown in Table 1). Supplementary Fig. 9 presents the distribution of sleep staging accuracy and Kappa across different datasets, indicating no significant differences in model performance between healthy subjects and patients with varying degrees of SDB. In reality, frequent abnormal events such as sleep apnea can lead to repeated micro-arousals and abnormal sleep structures. As the severity of SDB increases, sleep fragmentation worsens (as shown in Fig. 4i), making accurate sleep staging more challenging and placing higher demands on model performance. Previous studies15,45 have also confirmed this. However, our model’s performance did not decline with increasing SDB severity. This can be attributed to the following reasons: Firstly, we used a more extensive dataset, with a larger number of individuals with SDB included in the model training compared to previous studies; Additionally, the introduction of the auxiliary task enhanced the model’s ability to recognize sleep stages in patients with SDB.
Chronic comorbidities correlation analysis
Sleep is closely associated with chronic cardiovascular diseases46, respiratory diseases, and PD. These conditions can have an impact on sleep quality and alter respiratory signals, making accurate sleep staging more challenging. In this study, we evaluated the model’s performance in the context of three representative comorbidities involving the heart (hypertension), lungs (lung diseases), and brain (PD).
The results in Supplementary Fig. 10 show that the model performs similarly in subjects with and without hypertension. While hypertension may not directly alter respiratory characteristics, recent studies suggest that respiratory effort during sleep, commonly elevated in obstructive sleep apnea (OSA), may serve as an essential predictor of prevalent hypertension, potentially offering a novel biomarker for cardiovascular risk assessment. In contrast, the model’s performance is lower in subjects with lung diseases (Supplementary Fig. 11) than in healthy individuals. Lung diseases are expected to influence respiratory signals through changes in pulmonary mechanics and respiratory patterns, although the extent to which these changes reshape sleep architecture remains to be fully investigated. From another perspective, respiratory signals can serve as potential biomarkers for identifying lung diseases, suggesting the possibility of using respiratory signals for lung disease detection in future studies. Supplementary Fig. 12 illustrates the impact of PD on the model’s performance. The results indicate a significant decline in performance among PD subjects. However, due to the severe imbalance in the dataset’s ratio of PD to non-PD subjects, further in-depth research focusing specifically on individuals with PD is needed to accurately assess the model’s performance in this population. Additionally, in the external datasets, there is an issue of missing information regarding the discussed disease categories. In the future, it will be necessary to supplement the relevant data to further explore the relationship between sleep and comorbidities.
Real-time sleep staging
To extend the utility of our framework to online applications, we further evaluated its performance in real-time sleep staging via a self-supervised learning strategy (Fig. 2, Phase 3). In our study, “real-time” refers to the model’s ability to provide sleep stage predictions epoch-by-epoch, meaning that each 30-second segment is analyzed immediately after its completion, without access to future data. This real-time capability enables the early detection of potential sleep disorders, such as insomnia, and provides a foundation for timely and precise medical interventions.
We evaluated real-time sleep staging performance across different Historical Label Mapping Length (HLML) settings (1–5 minutes). Table 3 presents the results from the ClinHuaiAn and ClinRadar datasets, displaying statistical data such as accuracy by organizing all individuals’ predictions. From the perspective of model complexity, the model’s MFLOPs exhibit an approximately linear increase with the length of the input segments, which also leads to an increase in overall training time. In terms of sleep staging accuracy, the detection accuracy for 3, 4, and 5-minute input segments in the ClinHuaiAn dataset is higher than that for 1 or 2-minute segments. When the input length is extended from 1 minute to 5 minutes, the sleep staging accuracy improves by 1.29%, and the Kappa increases by 0.02. However, there is no significant performance difference between input lengths from 1 to 5 minutes, with an even smaller improvement observed in the ClinRadar dataset. These results indicate that the model performs consistently across different input segment lengths, and this self-supervised framework can achieve accurate real-time sleep stage prediction with data as short as 1 minute.
To further illustrate the temporal performance, Supplementary Fig. 13 compares the real-time sleep staging results with different input durations, pseudo-labels generated from full-night signal input, and PSG sleep labels. All of these methods effectively reflect the overall sleep trend throughout the night. When more surrounding sleep cycles are considered, the predictions from full-night input show smoother results with fewer frequent transitions. In contrast, the real-time results exhibit more frequent transitions due to the limitations of input information, relying only on data up to the current state. Consequently, balancing model complexity and detection accuracy is essential in practical applications, requiring an appropriate input segment length based on hardware capabilities. From the perspective of showcasing optimal results, and given the sufficient hardware capabilities in this study, we opted for 5 minutes as the input length for real-time sleep staging. To demonstrate the performance of real-time sleep staging, a representative example in a clinical environment is provided in Supplementary Movie 2.
Remote sleep management
Framework overview
We developed a remote sleep management platform, which facilitates efficient data transmission and user interaction, accommodating both historical analysis and real-time monitoring. As shown in Fig. 5, the system initiates with a radar sensor equipped with a Wi-Fi module that captures respiratory signals. These data are transmitted to a Message Queuing Telemetry Transport (MQTT) broker via the MQTT protocol, which serves as an intermediary, forwarding the data to a cloud server for further processing and storage. The cloud server, equipped with computational and storage capabilities, enables remote sleep health management. To ensure flexible access, the platform supports dual communication protocols. For accessing historical sleep data and analysis results, both clinician-facing and patient-facing applications communicate with the cloud server via Hypertext Transfer Protocol (HTTP) connections, allowing users to request and retrieve stored information. For real-time monitoring, the system establishes a WebSocket connection with the cloud server, enabling continuous updates and low-latency data transmission to clinicians and patient-facing interfaces.
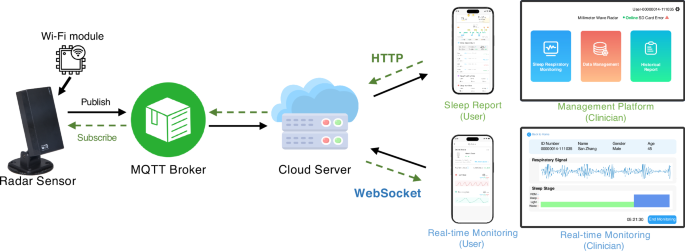
The diagram illustrates the end-to-end architecture of the platform, encompassing data collection, processing, and distribution. It highlights the key components, including radar sensors for non-contact data acquisition, the MQTT broker for efficient data transmission, the cloud server for computation and storage, and the user interfaces (clinician-facing and patient-facing) for accessing real-time and historical sleep health insights.
Model deployment
Our deployment process integrates TensorFlow-based models into the cloud server to support key analytical tasks such as sleep staging, AHI estimation, and real-time sleep staging. These models are containerized using Docker, enabling seamless deployment and scalability within the cloud environment. The deployed models subscribe to incoming data streams from the MQTT broker, process the data in real-time, and store the results in the server’s database. To address the challenge of high concurrency, a load balancer is employed to distribute incoming requests across multiple server instances. This approach supports horizontal scaling, allowing the system to handle increased traffic by dynamically adding more server instances as needed, ensuring stability and efficiency during peak loads. This deployment strategy ensures efficient, accurate, and continuous operation, enabling the platform to deliver timely and reliable sleep health insights while allowing for easy updates and optimizations to the models.
Representative case studies
In this part, we demonstrated the platform’s capability to effectively detect, manage, and evaluate treatment outcomes through representative case studies.
Case of insomnia
A female patient with a primary complaint of poor sleep quality during the night is investigated in this case. Initial monitoring by our platform on February 17, 2023, (Fig. 6a) confirmed significant insomnia symptoms, characterized by fragmented sleep patterns and reduced deep sleep and REM sleep. Based on the monitoring results, the platform recommended that the patient seek medical consultation. Following her visit to the hospital, the physician prescribed medication. Subsequent monitoring on February 21, 2023, revealed noticeable improvement, with reduced wakefulness and increased deep sleep and REM stages. After continuing the treatment, monitoring on April 06, 2023, showed significant improvement, with a more consolidated sleep structure. In light of this progress, the physician recommended reducing the medication dosage. The summary chart in Fig. 6a highlights the statistical changes across these stages, confirming the effectiveness of the treatment in improving sleep quality. These results underscore the platform’s ability to identify sleep issues and provide continuous feedback on treatment effectiveness, enabling personalized adjustments to the patient’s treatment plan.
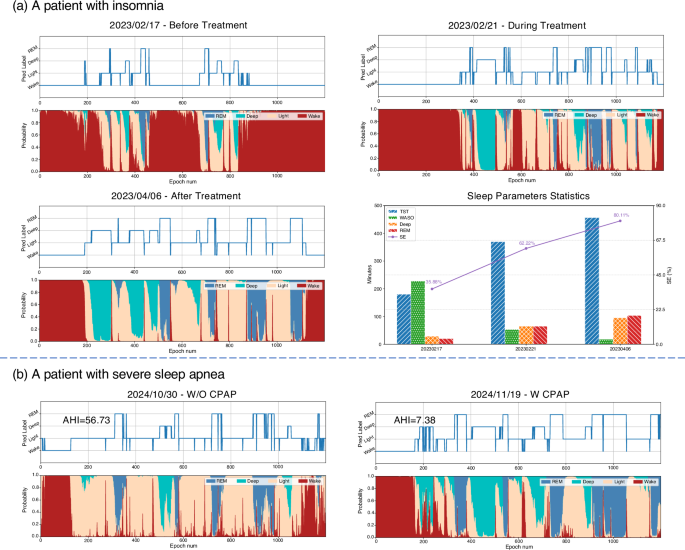
a A patient with insomnia: Sleep stage predictions and parameters (TST, SE, WASO, Deep sleep duration, and REM sleep duration) change for an insomnia patient at three time points: before treatment, during treatment, and after treatment, showing progressive improvements. Source data are provided as a Source Data file. b A patient with severe sleep apnea: Sleep stage predictions and AHI changes for a severe sleep apnea patient before and during CPAP treatment, demonstrating significant improvements in sleep structure and respiratory events.
Case of severe sleep apnea
In another demonstration, our platform effectively identifies and manages severe sleep disorders, as demonstrated by the case of a male patient from a remote area (Fig. 6b). Using our deployed non-contact radar device, the system detected severe sleep apnea symptoms and highly irregular sleep rhythms on October 30, 2024, with an initial AHI of 56.73, indicating a significant sleep disorder. Based on the monitoring data, the platform recommended immediate medical attention. The patient traveled to the sleep center of a Grade-A tertiary hospital in the city, where he was diagnosed with severe SDB. Physicians prescribed continuous positive airway pressure (CPAP) therapy. After CPAP treatment at home, the patient’s sleep quality improved dramatically. Monitoring on November 19, 2024, revealed a markedly reduced AHI of 7.38 and significant improvement in sleep architecture, including increased deep sleep and REM stages. This case highlights the platform’s ability to detect severe sleep disorders early, enabling timely intervention and supporting effective treatment outcomes through continuous monitoring.
These case studies highlight the platform’s capability for precise detection, personalized management, and comprehensive evaluation of treatment outcomes. In the future, the platform aims to further expand its functionality in the management of other sleep-related chronic diseases while enhancing its real-time analysis capabilities. Ultimately, with this platform, we aim to realize improving sleep health for the general population and reduce disparities in global sleep healthcare equity.
link